In this instalment we'll look at some more of the Hadoop-specific jobs.
Correlation/covariance matrix creation job
This Hadoop job produces either a correlation or covariance matrix (in a format that Weka's Matrix class can parse). It can handle numeric or nominal data (or a mixture of both) but the correlation matrix is only computed from the numeric fields in the data. Like the other remaining Hadoop jobs in the distributedWekaHadoop package, it requires that the ARFF header job be run first, and will run it automatically for you unless the -use-existing-header option is specified. The correlation matrix job relies on the summary meta data attributes computed by the ARFF header job so that it can compute the matrix in one pass over the data.
As explained in the first instalment, the map tasks compute a partial matrix of covariance sums for all rows in the matrix using their chunk of the data. The reducer phase aggregates on the basis of individual rows in the matrix, so the maps output rows of the full, but partially computed, matrix one at a time. This allows plenty of parallelism in the reduce phase, but does create lots of final output files (one for each reducer) that contain some of the rows of the final matrix. The job automatically tidies this up by reading all the part-r-xxxxx files and writing back to HDFS the final matrix in Weka's textual format. The job has an option to specify how many nodes are in the user's cluster, and then sets the number of reducers to min(num nodes * max reducers to run in parallel per node, num rows in the matrix).
The job also has options to specify a class attribute (which, if it happens to be a numeric field, is not part of the correlation analysis unless "keepClassAttributeIfSet" is selected) and run a principal components analysis in a post-processing phase after the job completes. The later does not distribute the computation of PCA - it runs locally on the client machine and the results are written back into the HDFS output directory. As PCA has a runtime that is at best quadratic in the number of input fields, this phase of the job is suitable for datasets that don't have tons of attributes. The PCA phase also creates a special serialised Weka filter that can be used for pre-processing in the classifier job.
Weka classifier builder job
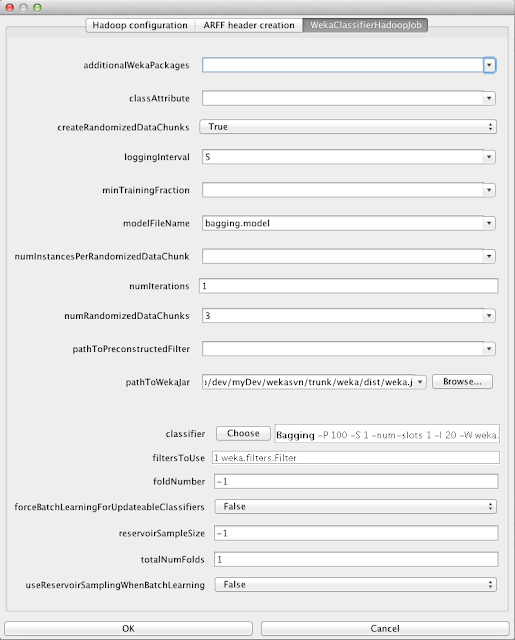
This job uses map-reduce to build classifier models in Hadoop and is one of the most complicated due to the number of options it provides. It will run up to three distinct jobs types depending on options:
- ARFF header creation (can be omitted if this has already run previously)
- Optional creation of randomly shuffled (and stratified) input data chunks from the original data
- Training of a Weka model (can involve multiple passes/jobs over the entire data set in the case of iterative incremental algorithms like SGD)
Note that separate runs of the randomised chunk creation phase may not be deterministic (even with the same random seed) due to the fact that keys (there is one per chunk) output from the map tasks are not guaranteed to arrive at the reducer in the same order from run to run, combined with the way the reducer "deals" instances out to the output files. However, once run, the randomly shuffled chunks may be re-used in subsequent model building and evaluation tasks.
The model learning phase handles a number of different training scenarios:
- Aggregateable classifiers (produce one final model of the same type as the individual models)
- Incremental aggregateable classifiers - e.g. naive Bayes, naive Bayes multinomial and SGD
- Batch aggregateable classifiers - e.g. Bagging
- Non-aggregateable classifiers
- Incremental ones - e.g. Hoeffding trees, raced incremental logit boost
- Batch learners - the majority of the classifiers and regressors in Weka
In the case of non-aggregateable classifiers, the final model produced in the reduce phase is a voted ensemble of the models learned by the mappers. For the technically oriented, this is essentially a "Dagging" model. In all cases, the final serialised model is deposited in a "model" subdirectory of the output path of the job in HDFS, along with a copy of the ARFF header (sans summary attributes). The header can be prepended to new data sets and makes the model ready for deployment wherever required.
A note on Bagging: The job makes a special check for Bagging (actually for any method that extends weka.classifiers.IteratedSingleClassifierEnhancer) and will divide the total requested number of base models by the number of map tasks that will be run. Thus Bagging runs in each mapper in order to produce some of the total number of base models requested by the user. The random forest algorithm can be implemented by setting the base learner to RandomTree in Bagging. I guess the final model produced by using Bagging in Hadoop in this fashion is actually a Dagging one again, with the small difference that the base models trained by each map will have training datasets created by bootstrap sampling on the data chunk that enters the map.
The classifier job has a number of options to fine tune the creation of the final model:
- If not using the option to create randomly shuffled data chunks, and instead the mapredMaxSplitSize option is used to control the number of maps, then minTrainingFraction can be used to prune away a model created on a data split that contains less data that the others
- numIterations controls how many times the model learning phase is invoked. This option only makes sense for iterative incremental classifiers such as SGD. Each iteration is a separate complete pass over the data. Hadoop's distributed cache is used to distribute the intermediate model learned at iteration i out to the nodes for training to continue in the mappers at iteration i + 1
- pathToPreconstructedFilter allows the PCA filter optionally created by the correlation matrix job to be used to transform the data coming into each map
- filtersToUse can be used instead of (or in conjunction with) pathToPreconstructedFilter in order to use standard Weka filters to pre-process data entering a map. The job automatically determines whether a given filter can be used with the base classifier and wraps the base classifier in one of several special subclasses of Weka's FilteredClassifier. For example, in order to maintain the ability to aggregate an Aggregateable classifier after filtering the input data, it is necessary that all filters used with it are Streamable - i.e., they can determine their output format from only header information (so that the output format produced by the filter is the same in each map) and don't buffer input data
- forceBatchLearningForUpdateableClassifiers will, as the name suggests, force an incremental classifier to be trained in a batch fashion. This essentially makes no difference for naive Bayes but does for SGD, which will perform a number of epochs over the training data entering a given map when trained in a batch fashion.
- useReservoirSamplingWhenBatchLearning results in the data streamed into each map getting passed into a reservoir sampling filter. This is useful to control the total number of instances processed in batch learning when the user is too lazy to tune the number of maps carefully via the mapredMaxSplitSize option or the option to create randomly shuffled data chunks.
Hmm. I thought I'd be able to cover everything in two instalments of this series of blog postings. Looks like I'll need a third. To be continued...





Hi,
ReplyDeleteThis post has been very helpful, Thanks :)
I would like to know how to omit the ARFFHeader creation job from the WekaClassifierHadoopJob in the KnowlwdgeFlow environment, I am unable to find any control on the frontend for doin the same.
Thanks in Advance.
On the "ARFF header creation" tab enter a path to a header file in the "pathToExistingHeader" field. This can be a local file or the output of a previous run of the ARFF header job in HDFS.
ReplyDeleteCheers,
Mark.
Thanks for the reply.
ReplyDeleteHow do i associate the output of the previous task to the next one.
i.e. provide the hdfs path of the ARFF header generated in previous run
You will have to fill this field in manually. However, if you specify the name of the ARFF header to create in the ARFF header job ("outputHeaderFileName" field), then the full path is just the output path of the ARFF header job + that file name. This can be set in the "pathToExistingHeader" field in other jobs in advance as long as the ARFF header job runs first of course.
ReplyDeleteCheers,
Mark.
Thanks again.
ReplyDeleteWhen i run multiple jobs i.e. a classifier job followed by a scoring job , the jars are loaded in hdfs for each job . so is there a way to restrict the loading of jars to one job and the consecutive jobs refering to the jars installed by the previous one??
Not at present I'm afraid. I might add an option to prevent the installation of the jars into HDFS or perhaps just check to see if they already exist. Luckily there are not many jars and they are not very big, so the installation just takes a matter of seconds.
ReplyDeleteThe current behaviour is quite convenient for me while developing as I do lots of change + test iterations :-)
Cheers,
Mark.
Hi,
ReplyDeleteI tried running the ArffHeaderJob through Hadoop commandline.The arff directory is generated but the arff header file is not generated.The arff directory is empty.
Plz Help!
Are there any exceptions/errors on the command line, or in the Hadoop logs?
ReplyDeleteHELLO Sir,
ReplyDeleteI am trying to work with Neural Network in weka but it is not supporting to classify data with above 20 attributes and approx 10,000 records plz reply what solution shold I follow
Hello Mark,
ReplyDeleteCan I create Bayesian Networks in distributedWekaHadoop ?
hi sir i did nt get hdfs saver and hdfs loader in knowledge flow...can u help m out i have installed everything
ReplyDeleteThey appear under DataSources and DataSinks rather than under the Hadoop folder.
ReplyDeleteCheers,
Mark.
This comment has been removed by the author.
ReplyDeleteHello Mark
ReplyDeleteI run the WekaClassifierHadoopJob with a CSV file in inputPath and it worked and generates this files:
outputModel.model
outputModel_arffHeader.arff
-What is this outputModel.model and where I`ll use this file?
-Why the arff file did not come with complete data? Is it because the compression option?
Thanks
Hello Mark
ReplyDeleteI run the commands and aparently it is working. But something is strange in the cluster:
I Run a java toy program, without using the DistributedWekaHadoop, and I have the output with many files: part-r-00000, part-r-00001, part-r-00002 and part-r-00003. (One for each node in the cluster)
But, when I run the DistributedWekaHadoop job, I have the output with just one file: part-r-00000. So, now I don't know if the cluster is really working. How can I know If all datanodes in the cluster is working properly with the DistributedWekaHadoop?
*I have sure that I'm using the hadoop user in the cluster and the masternode and all datanodes are turned on.
Thank you
Many of the Weka jobs run only a single reducer (e.g. creating a model requires that all the partial models learned by the map tasks are reduced by one reducer into a single final model), this is why there is only one part-r file output. The number of mappers run is where the parallelism is happening, and this is typically controlled by the file block size (as one map task is used for each block).
ReplyDeleteCheers,
Mark.
Hello Mark
ReplyDeleteI'm trying to run the BayesNet algorithm using ADTrees on the cluster, but I'm with this problem: "Not enough memory (less than 50MB left on heap). Please load a smaller dataset or use a larger heap size". I have the same error with a single computer, but with the cluster I expected that would be possible to run.
I Already tried to use the command: java -Xmx512m -jar weka.jar to run the Weka, but did not work. How can I fix this problem?
Thanks
This sounds like an exception that is generated by Weka's GUI, in which case it is not an error occurring out on the cluster. It could be that the model generated by ADTree in the cluster (which is pulled back to the Weka client) is too large for the available heap memory in the client JVM. 512Mb is pretty small these days, try using 1Gb or more.
ReplyDeleteCheers,
Mark.
Hello Mark
DeleteI already added the command -Xmx3g (my masternode RAM memory has 4g) and this works in many situations, but now I need to use an even more advanced configuration and I'm facing this problem again. What more else can I do to solve this problem?
Thanks
Hello Mark
ReplyDeleteI would like to know how can I save a BifFile, using the commands in the cluster? I need to use the Bif XML format to compare with another bayes networks.
I found the -d command at the documentation, but it did not work in the cluster. I try to use the command: -model-file-name Bayes.xml (because if I save with .model, it is not possible to use in the Biffile), but the results was the same.
I found another documentation with this information: "How do I save a Bayes net in BIF format? Command line: use the-g option and redirect the output on stdout into a File". So, I already try to use the command: -W weka.classifiers.bayes.BayesNet -g testando.xml --... but it did not work.
So, What are the commands to save a BifFile using the command line in the cluster?
Thanks
I think you are out of luck here. The main reason is that the final model built in the cluster is not a single Bayes net. Instead, it is a voted ensemble of Bayes nets (one net learned by each mapper that runs on a block of your data).
ReplyDeleteThe main aim of Weka's distributed support was to enable any classifier to be learned in the cluster in some fashion. For embarrassingly parallel models (i.e. naive Bayes variants and linear models combined via averaging) you get a single model of the same type produced. For all other classifiers the voted ensemble approach is used.
Cheers,
Mark.
Hello Mark
DeleteOk, but Is it possible to generate a bayes network on a single computer and compare with a model generated on the cluster?
I try to do this using the commands: ... -W weka.classifiers.bayes.BayesNet -- -D -B /user/net.xml ... but the comparison did not work as on the single computer where I get this results for the comparison:
Missing: 3 Extra: 31 Reversed: 0
Divergence: -1.7779137843688664
The result file on the cluster only contains the normal results, with no comparison metrics. Is it not possible to do on distributed Weka or am I using the incorrect commands?
Thanks
Hello Mark
ReplyDeleteI'm reading and trying to understand the voted ensemble for combining multiple models process. I see that exists three schemes to combine multiple models: bagging, boosting and stacking.
So, It seems that when I run Weka on a single computer, some classifiers algorithms can also uses the voting method. If I understand correctly, why this voted ensemble process does not allow to generate a biffile on the cluster If it is possible to generate on a single computer?
I try to use the same commands that worked perfectly on the single computer (to generate a biffile and after, to compare with new bayes networks using another algorithms), but it's not working on the cluster (as I commented before at your last answer). Can you help me to understand, please?
Thanks
In the case of distributed Weka, where a model is trained on each part of the full dataset (i.e. a block of the input data as defined by the Hadoop file system, or a partition as defined by a Spark RDD) a voted ensemble just combines each model created from each block/partition. This is effectively a "Dagged" (Disjoint Aggregation) model; not to be confused with a Bagged (Bootstrap Aggregation) model.
ReplyDeleteThe point here is that there are multiple Bayes nets, potentially with different network structures. How do you propose to produce one XML Biff file to describe this? It would be possible to add some functionality to output the XML for each of the base models individually. Note that the Vote meta classifier does not implement Weka's Drawable interface, so even on a single machine it is not possible to output a graph from this classifier.
Cheers,
Mark.
Hello Mark
DeleteOk, great!
Thank you
Dear Mark
ReplyDeleteGreat job
Kindly, can you provide some examples on part 2 and 3 ..
Nice Article Very Helpful ! Thanks for sharing ! Also check
ReplyDeleteAttitude Shayari
Sad Hindi Shayari
Shayari99
Thanks for you for adding all of them in one place! it is difficult to get the throw right with therefore many heroes! and with the reveals being just once weekly, by enough time you receive them down in your head, you do not see them again! Congrats, and thank you for remaining correct to the collection and perhaps not giving spoilers! I'm amazingly satisfied with the way the series sticks to the specific discussion of the publications! And I think the castings have already been amazing! I specially like Tyreon...his identity has more school than all others of his household!New Celebrity Net Worth Gucci Mane
ReplyDeleteNew Celebrity Net Worth Eminem
New Celebrity Net Beyonce
New Celebrity Net Adam Sandler
New Celebrity Net Jerry Seinfeld
New Celebrity Net Kevin Hart
New Celebrity Net Triple H
New Celebrity Net John Cena
New Celebrity Net Julia Louis Dreyfus
New Celebrity Net David Bowie
This was really helpful
ReplyDeleteThis comment has been removed by the author.
ReplyDeleteGreat Post, very helpfull for me...
ReplyDeletethesrvtech
Whatsapp Me Dark Mode Kaise Enable kare
Podcast Kya hai
Google Meet App Kya Hai
Paytm Service Agent kaise bane
Reselling Kya Hai
hello friends my name is faug Faug Game Download Apk download this game for your android mobile
ReplyDeleteFaug Game Download Apk
love it!!! Great job
ReplyDeleteHP Printer Support
thanks for giving that type of information. Really enjoyed this blog post. Really looking forward to reading more. Much obliged.
Top Recruitment Consultancy
It is in detail and well formatted that i enjoyed reading. which inturn helped me to get new information from your blog.
Contact Folks - All in One Helpline
This content creates a new hope and inspiration with in me.
ReplyDeleteBest Sorry Status in Hindi for Whatsapp and Facebook of 2021
sorry status
sorry status
sorry status
sorry status
sorry status
au-usd Is The World's First Exchange That Allows You To Trade digital Currencies In Real-Time Using The Power Of Artificial Intelligence.
ReplyDeletenice nice
ReplyDeleteInternship providing companies in chennai | Where to do internship | internship opportunities in Chennai | internship offer letter | What internship should i do | How internship works | how many internships should i do ? | internship and inplant training difference | internship guidelines for students | why internship is necessary
This comment has been removed by the author.
ReplyDeleteDo you believe in long term investement . One of the option of doing investement is by investing in Crypto currencies. You can invest in Fudxcoin company that deals in the selling and purchasing of Crypto Currency. It is a reliable company. One need not doubt in investing in it as i have also bought crypto currency from it and feeling very satisfied with their services.
ReplyDeletecrypto currency block chain technology
I want the world to know about where to invest their hard earned money and get fruitful returns. If one is looking forward of investing he can go into investment of crypto coins.
You can invest in Fudxcoin company that deals in the selling and purchasing of Crypto Currency. It is a reliable company. One need not doubt in investing in it as i have also bought crypto currency from it and feeling very satisfied with their services.
crypto currency block chain technology